This is Part 2 of the “RAG is all you need” series. If you haven’t read Part 1 where we covered how LLMs work and why grounding them to known truth matters, start there.
Introduction
Last time we talked about why RAG is becoming essential for AI applications, right? It’s kind of one of those tools that every AI engineer ends up reaching for.
Today we’re going after vector search. And this is interesting because once you get this concept, a lot of things about modern AI just click.
There’s a question that’s been bugging computer scientists for ages: how do we get computers to understand language the way we do? Try this. Think about these words:
- table
- flower
- lamp
- basketball
If I throw in the word “chair” and ask you which one’s most related, you’d probably say table.
But if I asked you why, you might start talking about how chairs and tables go together during meals, or how they’re both made of wood. But basketball boards are made of wood too. And you sit near lamps all the time. See how messy this gets?
NLP researchers have been grinding away at better ways to handle these nuanced relationships. They started with stuff like bag-of-words (kinda naive, if you ask me), but then BERT came along and changed the whole picture.
What BERT and similar models do is map words into a dense vector space where relationships are represented by how words appear together in massive amounts of text. They’re building this multidimensional map of language itself.
And when I say multidimensional, I mean way more than the 2D diagram above. Think 512 dimensions or more.
The way I like to think about it: every word gets its own unique “location” in this space, and words that often show up together in similar contexts end up closer to each other. Modern language models like Claude and GPT have gotten remarkably good at this mapping.
Understanding Vector Embeddings in Practice
When it comes to actually building AI applications (isn’t that why we’re all here?), we need to take our knowledge base, our ground truth, and convert it into vector embeddings. This is where you start to differentiate your AI chatbot from everyone else’s. The uniqueness and usefulness of the data you bolt on is what matters: company documentation, school syllabi, performance reports, pretty much any text you want your AI to understand.
There’s several approaches you could take for creating these embeddings. You could use smaller models like BERT, and honestly? If you’re just getting started or working on a smaller project, that’s totally fine. You can do it absolutely free with open-source BERT models.
But in production, the way I like to do it is reach for high-dimension, high-accuracy embedding models. I’m talking about models from OpenAI, for example. They just handle the tricky, ambiguous language tasks so much better.
Let’s build this up piece by piece. First, we need to pick an embedding model. LangChain makes this pretty painless:
from langchain.embeddings import OpenAIEmbeddings, HuggingFaceEmbeddings
# Option A: OpenAI - better accuracy, costs money
embeddings = OpenAIEmbeddings()
# Option B: Open-source BERT model - free, good enough for testing
embeddings = HuggingFaceEmbeddings(
model_name="sentence-transformers/all-MiniLM-L6-v2"
)
So that’s the first thing. You’ve got your embedding model wired up. But you can’t just throw an entire document at it. Most embedding models have a token limit (OpenAI’s text-embedding-3-small maxes out at 8191 tokens). So you need to break your text into chunks first.
Let me make this concrete. Say we’re building a RAG system for a Malawian secondary school, and we want to embed this biology textbook passage:
The human circulatory system is a complex network of blood vessels, the heart, and blood. The heart is a muscular organ roughly the size of a closed fist. It pumps blood through the body via two circuits: the pulmonary circuit (heart to lungs and back) and the systemic circuit (heart to the rest of the body and back). Arteries carry oxygenated blood away from the heart, while veins return deoxygenated blood. Capillaries are the smallest vessels, connecting arteries to veins, and this is where gas exchange actually happens at the tissue level. The average adult heart beats about 72 times per minute, pumping roughly 5 litres of blood through 96,000 kilometres of blood vessels.
That’s about 120 words. A real textbook chapter might be 5,000+ words. You can’t embed the whole thing at once, and even if you could, the embedding would be too diluted to match specific questions like “what do capillaries do?”
This is where RecursiveCharacterTextSplitter comes in:
from langchain.text_splitter import RecursiveCharacterTextSplitter
text_splitter = RecursiveCharacterTextSplitter(
chunk_size=150, # characters per chunk (small for demo)
chunk_overlap=30, # characters of overlap between chunks
length_function=len,
)
What chunking actually produces
Using our biology text with chunk_size=150 and chunk_overlap=30, here’s what the splitter gives us:
CHUNK 1 (chars 0-149):
"The human circulatory system is a complex network of blood
vessels, the heart, and blood. The heart is a muscular organ
roughly the size of a closed fist."
CHUNK 2 (chars 120-269):
"the size of a closed fist. It pumps blood through the body
via two circuits: the pulmonary circuit (heart to lungs and
back) and the systemic circuit"
↑
overlap: "the size of a closed fist." appears in both chunks
CHUNK 3 (chars 240-389):
"and the systemic circuit (heart to the rest of the body and
back). Arteries carry oxygenated blood away from the heart,
while veins return deoxygenated blood."
↑
overlap: "and the systemic circuit" bridges the two chunks
See what’s happening? Each chunk has a 30-character overlap with the previous one. That overlap is load-bearing. Without it, Chunk 2 would start mid-sentence with “It pumps blood through the body” and the model would have no idea what “it” refers to.
What happens without overlap
Let’s see the same text with chunk_overlap=0:
CHUNK 1: "...The heart is a muscular organ roughly the size of a"
CHUNK 2: "closed fist. It pumps blood through the body via two..."
“a closed fist” got sliced in half. If someone asks “what size is the heart?”, neither chunk has a complete answer. The overlap prevents this by making sure sentences at the boundary appear in both chunks.
In production, I typically use chunk_size=1000 with chunk_overlap=200. That gives you roughly paragraph-sized chunks with enough overlap to preserve context at the edges.
# Production settings
text_splitter = RecursiveCharacterTextSplitter(
chunk_size=1000,
chunk_overlap=200,
length_function=len,
)
chunks = text_splitter.split_text(biology_chapter)
# A 5,000-char chapter produces roughly 6 chunks
From chunks to vectors
Now we wire the splitter to the embedding model. For each chunk, we generate a vector and store it with metadata so we can trace it back to the source:
from typing import List, Dict
async def generate_embeddings(
text: str, embeddings, text_splitter
) -> List[Dict]:
chunks = text_splitter.split_text(text)
results = []
for i, chunk in enumerate(chunks):
vector = await embeddings.aembed_query(chunk)
results.append({
"text": chunk,
"vector": vector, # e.g. 1536 floats for OpenAI
"metadata": {
"chunk_index": i,
"length": len(chunk),
"source": "biology_ch3", # trace back to the document
}
})
return results
And to see it end to end:
async def main():
embeddings = OpenAIEmbeddings()
text_splitter = RecursiveCharacterTextSplitter(
chunk_size=1000, chunk_overlap=200, length_function=len
)
# Our biology textbook chapter
chapter_text = load_chapter("biology_ch3.txt")
results = await generate_embeddings(
chapter_text, embeddings, text_splitter
)
print(f"Generated {len(results)} chunks")
print(f"Vector size: {len(results[0]['vector'])} dimensions")
# Output: Generated 6 chunks
# Output: Vector size: 1536 dimensions
So we went from a raw textbook chapter to a list of 1536-dimension vectors, each traceable back to a specific paragraph. When a student later asks “what connects arteries to veins?”, the vector search will surface Chunk 3 (the one about capillaries and gas exchange) because that question and that chunk will be close neighbours in vector space.
That’s the whole pipeline in about 30 lines.
The Vector Database Problem
You and I know all about Relational Database Management Systems (RDBMSs), right? They’re built for storing stuff in rows and columns. Very 2D thinking. Some databases get fancier with graphs or nodes (looking at you, Neo4j). But vector databases are purpose-built for a different problem entirely.
They can handle information in vector space that’s hundreds or even thousands of dimensions. And over the last few years, we’ve seen some solid options emerge:
- Pinecone (built specifically for vector search)
- Chroma (built specifically for vector search)
- PostgreSQL with pg_vector (for when you want to stick with the classics)
- Redis (my personal favourite, because performance)
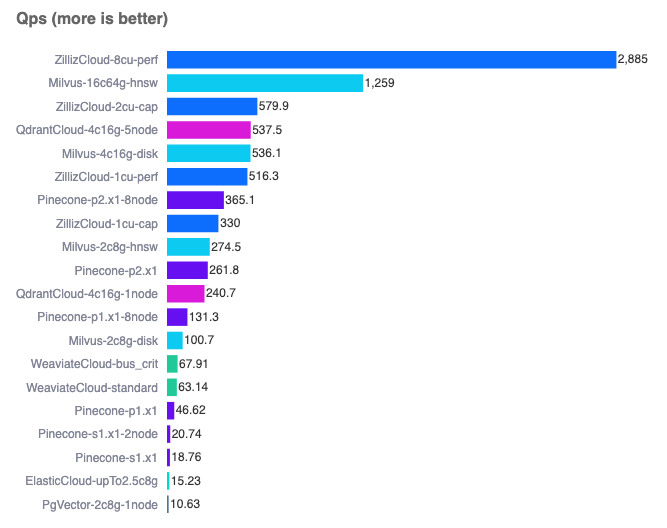
You can run a comparison yourself here.
Building The Pipeline
So here’s how the whole thing actually works in practice. When someone asks a question, we:
- Store your ground truth in a vector database
- Convert the user’s question into an embedding
- Use cosine similarity (actually a pretty simple algorithm) to find matches
- Pull back maybe the top 10 most relevant chunks of information
There’s a whole science to storing these embeddings well. It depends on which LLM you’re using, the context window size you’re working with, and how much data you can throw at it before things get wonky.
The good news? Newer models with their huge context windows have sanded down a lot of the rough edges here. You can throw a pretty big chunk of context at them and they handle it well.
The hallucination problem hasn’t disappeared entirely (probably never will), but it’s gotten a lot more manageable.
Practical Implementation Tips
When you’re actually implementing this, you want to:
- Chunk your data into manageable sizes
- Store references to your original documents
- Fetch those reference docs when you get a match
This way, when your LLM is answering questions, it’s pulling from actual, accurate information from your knowledge base. Not just its training data.
Below is a representation of how you could store your RAG data using Postgres:
documents
| Column | Type | Description |
|---|---|---|
| id | UUID | Primary key, unique document identifier |
| title | TEXT | Document title/name |
| source_url | TEXT | Where this doc came from (optional) |
| created_at | TIMESTAMP WITH TIMEZONE | When we first added this |
| updated_at | TIMESTAMP WITH TIMEZONE | Last time we touched this |
document_chunks
| Column | Type | Description |
|---|---|---|
| id | UUID | Primary key, unique chunk identifier |
| document_id | UUID | Links back to our main document |
| chunk_index | INTEGER | Keeps chunks in order (important) |
| content | TEXT | The actual chunk of text |
| embedding | vector(1536) | The vector embedding |
| metadata | JSONB | Flexible metadata storage |
| tokens | INTEGER | Token count (helpful for context windows) |
| chunk_size | INTEGER | Size of this chunk in chars |
| created_at | TIMESTAMP WITH TIMEZONE | When we created this chunk |
You know what’s cool about this setup? The metadata JSONB field is this flexible catch-all for whatever extra info you need to track. It becomes helpful for improving the relevance and accuracy of your search (more on that later in the series).
I usually stuff things in there like:
{
"source_type": "technical_doc",
"department": "engineering",
"last_verified": "2024-01-18",
"confidence": 0.95,
"page_number": 42,
"section": "API Documentation"
}
Next up, we’ll roll up our sleeves and implement a real world RAG solution against the MANEB syllabus, which can be used later to build a context-aware AI tutor for Malawian students.
